Federated Kmeans¶
Kmeans is a simple statistic model widely used for clustering. FATE provides Heterogeneous Kmeans(HeteroKmeans).
Here we simplify participants of the federation process into three parties. Party A represents Guest, party B represents Host. Party C, which is also known as “Arbiter,” is a third party that works as a coordinator. Party C is responsible for generating private and public keys.
Heterogeneous Kmeans¶
The process of HeteroKmeans training is shown below:
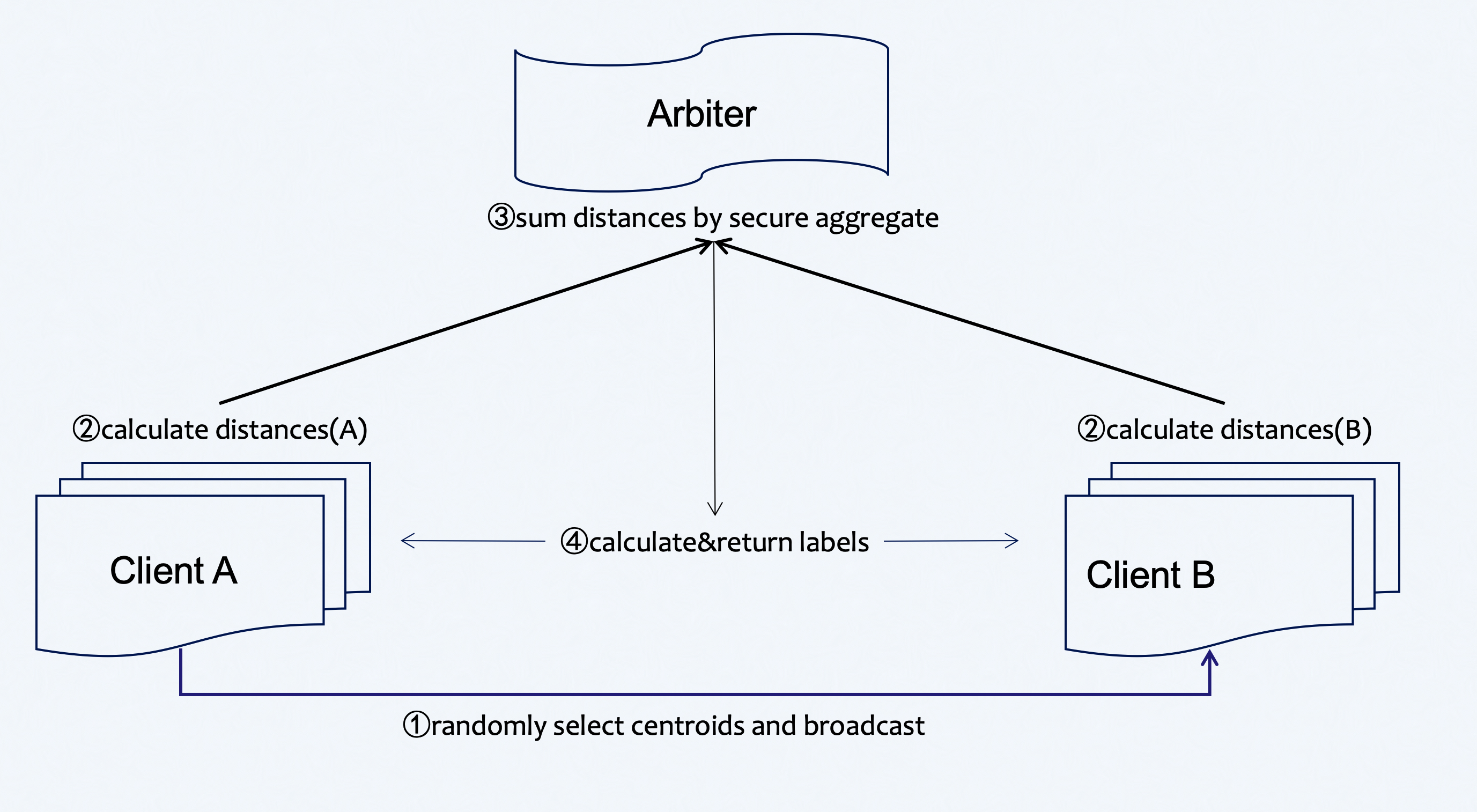
Figure 1 (Federated HeteroKmeans Principle)
A sample alignment process is conducted before training. The sample alignment process identifies overlapping samples in databases of all parties. The federated model is built based on the overlapping samples. The whole sample alignment process is conducted in encryption mode, and so confidential information (e.g. sample ids) will not be leaked.
In the training process, party A will choose centroids from samples randomly and send them to party B . Party A and party B then compute the distance to centroids ,which is needed for label assignment. Arbiter aggregates, calculates, and returns back the final label to each sample and repeats this part until the max iter or tolerance meets the criteria.
During the aggregate process, parties will use secure aggregate as all sent distances will be added with random numbers that can be combined to zero when aggregating at arbiter.
Param¶
hetero_kmeans_param
¶
Classes¶
KmeansParam(k=5, max_iter=300, tol=0.001, random_stat=None)
¶
Bases: BaseParam
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
k |
int, default 5
|
The number of the centroids to generate. should be larger than 1 and less than 100 in this version |
5
|
max_iter |
int, default 300.
|
Maximum number of iterations of the hetero-k-means algorithm to run. |
300
|
tol |
float, default 0.001.
|
tol |
0.001
|
random_stat |
None or int
|
random seed |
None
|
Source code in python/federatedml/param/hetero_kmeans_param.py
38 39 40 41 42 43 | |
Attributes¶
k = k
instance-attribute
¶max_iter = max_iter
instance-attribute
¶tol = tol
instance-attribute
¶random_stat = random_stat
instance-attribute
¶Functions¶
check()
¶Source code in python/federatedml/param/hetero_kmeans_param.py
45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 | |
Features¶
- Tolerance & Max_iter supported for convergence
- Random_stat specify supported
- Centroids are selected randomly
- Labeled and unlabeled data supported
Examples¶
Example
## Hetero Kmeans Pipeline Example Usage Guide.
#### Example Tasks
This section introduces the Pipeline scripts for different types of tasks.
1. Train with Feature-engineering Task :
script: pipeline-kmeans-with-feature-enginnering.py
2. Multi-host Task:
script: pipeline-kmeans-multi-host.py
3. Train Task:
script: pipeline-kmeans.py
4. Train with validate Task:
script: pipeline-kmeans-validate.py
Users can run a pipeline job directly:
python ${pipeline_script}
pipeline-kmeans-multi-host.py
import argparse
from pipeline.backend.pipeline import PipeLine
from pipeline.component import DataTransform
from pipeline.component import HeteroKmeans
from pipeline.component import Intersection
from pipeline.component import Evaluation
from pipeline.component import Reader
from pipeline.interface import Data
from pipeline.utils.tools import load_job_config
def main(config="../../config.yaml", namespace=""):
if isinstance(config, str):
config = load_job_config(config)
parties = config.parties
guest = parties.guest[0]
hosts = parties.host
arbiter = parties.arbiter[0]
guest_train_data = {"name": "breast_hetero_guest", "namespace": f"experiment{namespace}"}
host_train_data = [{"name": "breast_hetero_host", "namespace": f"experiment{namespace}"},
{"name": "breast_hetero_host", "namespace": f"experiment{namespace}"}]
# initialize pipeline
pipeline = PipeLine()
# set job initiator
pipeline.set_initiator(role='guest', party_id=guest)
# set participants information
pipeline.set_roles(guest=guest, host=hosts, arbiter=arbiter)
# define Reader components to read in data
reader_0 = Reader(name="reader_0")
# configure Reader for guest
reader_0.get_party_instance(role='guest', party_id=guest).component_param(table=guest_train_data)
# configure Reader for host
reader_0.get_party_instance(role='host', party_id=hosts[0]).component_param(table=host_train_data[0])
reader_0.get_party_instance(role='host', party_id=hosts[1]).component_param(table=host_train_data[1])
# define DataTransform components
data_transform_0 = DataTransform(name="data_transform_0") # start component numbering at 0
# get DataTransform party instance of guest
data_transform_0_guest_party_instance = data_transform_0.get_party_instance(role='guest', party_id=guest)
# configure DataTransform for guest
data_transform_0_guest_party_instance.component_param(with_label=True, output_format="dense")
# get and configure DataTransform party instance of host
data_transform_0.get_party_instance(role='host', party_id=hosts[0]).component_param(with_label=False)
data_transform_0.get_party_instance(role='host', party_id=hosts[1]).component_param(with_label=False)
# define Intersection components
intersection_0 = Intersection(name="intersection_0")
param = {
"k": 3,
"max_iter": 10
}
hetero_kmeans_0 = HeteroKmeans(name='hetero_kmeans_0', **param)
evaluation_0 = Evaluation(name='evaluation_0', eval_type='clustering')
# add components to pipeline, in order of task execution
pipeline.add_component(reader_0)
pipeline.add_component(data_transform_0, data=Data(data=reader_0.output.data))
# set data input sources of intersection components
pipeline.add_component(intersection_0, data=Data(data=data_transform_0.output.data))
pipeline.add_component(hetero_kmeans_0,
data=Data(train_data=intersection_0.output.data))
# print(f"data: {hetero_kmeans_0.output.data.data[0]}")
pipeline.add_component(evaluation_0, data=Data(data=hetero_kmeans_0.output.data.data[0]))
# compile pipeline once finished adding modules, this step will form conf and dsl files for running job
pipeline.compile()
# fit model
pipeline.fit()
# query component summary
print(pipeline.get_component("hetero_kmeans_0").get_summary())
if __name__ == "__main__":
parser = argparse.ArgumentParser("PIPELINE DEMO")
parser.add_argument("-config", type=str,
help="config file")
args = parser.parse_args()
if args.config is not None:
main(args.config)
else:
main()
hetero_kmeans_testsuite.json
{
"data": [
{
"file": "examples/data/breast_hetero_guest.csv",
"head": 1,
"partition": 16,
"table_name": "breast_hetero_guest",
"namespace": "experiment",
"role": "guest_0"
},
{
"file": "examples/data/breast_hetero_host.csv",
"head": 1,
"partition": 16,
"table_name": "breast_hetero_host",
"namespace": "experiment",
"role": "host_0"
},
{
"file": "examples/data/breast_hetero_host.csv",
"head": 1,
"partition": 16,
"table_name": "breast_hetero_host",
"namespace": "experiment",
"role": "host_1"
},
{
"file": "examples/data/vehicle_scale_hetero_guest.csv",
"head": 1,
"partition": 16,
"table_name": "vehicle_scale_hetero_guest",
"namespace": "experiment",
"role": "guest_0"
},
{
"file": "examples/data/vehicle_scale_hetero_host.csv",
"head": 1,
"partition": 16,
"table_name": "vehicle_scale_hetero_host",
"namespace": "experiment",
"role": "host_0"
}
],
"pipeline_tasks": {
"kmeans": {
"script": "pipeline-kmeans.py"
},
"kmeans_validate": {
"script": "pipeline-kmeans-validate.py"
},
"kmeans_feature_engineering": {
"script": "pipeline-kmeans-with-feature-engineering.py"
},
"kmeans_multi_host": {
"script": "pipeline-kmeans-multi-host.py"
}
}
}
pipeline-kmeans.py
import argparse
from pipeline.backend.pipeline import PipeLine
from pipeline.component import DataTransform
from pipeline.component import HeteroKmeans
from pipeline.component import Intersection
from pipeline.component import Evaluation
from pipeline.component import Reader
from pipeline.interface import Data
from pipeline.utils.tools import load_job_config
def main(config="../../config.yaml", namespace=""):
# obtain config
if isinstance(config, str):
config = load_job_config(config)
parties = config.parties
guest = parties.guest[0]
host = parties.host[0]
arbiter = parties.arbiter[0]
guest_train_data = {"name": "breast_hetero_guest", "namespace": f"experiment{namespace}"}
host_train_data = {"name": "breast_hetero_host", "namespace": f"experiment{namespace}"}
# initialize pipeline
pipeline = PipeLine()
# set job initiator
pipeline.set_initiator(role='guest', party_id=guest)
# set participants information
pipeline.set_roles(guest=guest, host=host, arbiter=arbiter)
# define Reader components to read in data
reader_0 = Reader(name="reader_0")
# configure Reader for guest
reader_0.get_party_instance(role='guest', party_id=guest).component_param(table=guest_train_data)
# configure Reader for host
reader_0.get_party_instance(role='host', party_id=host).component_param(table=host_train_data)
# define DataTransform components
data_transform_0 = DataTransform(name="data_transform_0") # start component numbering at 0
# get DataTransform party instance of guest
data_transform_0_guest_party_instance = data_transform_0.get_party_instance(role='guest', party_id=guest)
# configure DataTransform for guest
data_transform_0_guest_party_instance.component_param(with_label=True, output_format="dense")
# get and configure DataTransform party instance of host
data_transform_0.get_party_instance(role='host', party_id=host).component_param(with_label=False)
# define Intersection components
intersection_0 = Intersection(name="intersection_0")
param = {
"k": 3,
"max_iter": 10
}
hetero_kmeans_0 = HeteroKmeans(name='hetero_kmeans_0', **param)
evaluation_0 = Evaluation(name='evaluation_0', eval_type='clustering')
# add components to pipeline, in order of task execution
pipeline.add_component(reader_0)
pipeline.add_component(data_transform_0, data=Data(data=reader_0.output.data))
# set data input sources of intersection components
pipeline.add_component(intersection_0, data=Data(data=data_transform_0.output.data))
# set train & validate data of hetero_lr_0 component
pipeline.add_component(hetero_kmeans_0, data=Data(train_data=intersection_0.output.data))
print(f"data: {hetero_kmeans_0.output.data.data[0]}")
pipeline.add_component(evaluation_0, data=Data(data=hetero_kmeans_0.output.data.data[0]))
# compile pipeline once finished adding modules, this step will form conf and dsl files for running job
pipeline.compile()
# fit model
pipeline.fit()
# query component summary
print(pipeline.get_component("hetero_kmeans_0").get_summary())
if __name__ == "__main__":
parser = argparse.ArgumentParser("PIPELINE DEMO")
parser.add_argument("-config", type=str,
help="config file")
args = parser.parse_args()
if args.config is not None:
main(args.config)
else:
main()
pipeline-kmeans-with-feature-engineering.py
import argparse
from pipeline.backend.pipeline import PipeLine
from pipeline.component import DataTransform
from pipeline.component import HeteroKmeans
from pipeline.component import Intersection
from pipeline.component import HeteroFeatureBinning
from pipeline.component import HeteroFeatureSelection
from pipeline.component import Evaluation
from pipeline.component import Reader
from pipeline.interface import Data
from pipeline.interface import Model
from pipeline.utils.tools import load_job_config
def main(config="../../config.yaml", namespace=""):
# obtain config
if isinstance(config, str):
config = load_job_config(config)
parties = config.parties
guest = parties.guest[0]
host = parties.host[0]
arbiter = parties.arbiter[0]
guest_train_data = {"name": "breast_hetero_guest", "namespace": f"experiment{namespace}"}
host_train_data = {"name": "breast_hetero_host", "namespace": f"experiment{namespace}"}
# initialize pipeline
pipeline = PipeLine()
# set job initiator
pipeline.set_initiator(role='guest', party_id=guest)
# set participants information
pipeline.set_roles(guest=guest, host=host, arbiter=arbiter)
# define Reader components to read in data
reader_0 = Reader(name="reader_0")
# configure Reader for guest
reader_0.get_party_instance(role='guest', party_id=guest).component_param(table=guest_train_data)
# configure Reader for host
reader_0.get_party_instance(role='host', party_id=host).component_param(table=host_train_data)
# define DataTransform components
data_transform_0 = DataTransform(name="data_transform_0") # start component numbering at 0
# get DataTransform party instance of guest
data_transform_0_guest_party_instance = data_transform_0.get_party_instance(role='guest', party_id=guest)
# configure DataTransform for guest
data_transform_0_guest_party_instance.component_param(with_label=True, output_format="dense")
# get and configure DataTransform party instance of host
data_transform_0.get_party_instance(role='host', party_id=host).component_param(with_label=False)
# define Intersection components
intersection_0 = Intersection(name="intersection_0")
param = {
"name": 'hetero_feature_binning_0',
"method": 'optimal',
"optimal_binning_param": {
"metric_method": "iv"
},
"bin_indexes": -1
}
hetero_feature_binning_0 = HeteroFeatureBinning(**param)
param = {
"name": 'hetero_feature_selection_0',
"filter_methods": ["manually", "iv_filter"],
"manually_param": {
"filter_out_indexes": [1]
},
"iv_param": {
"metrics": ["iv", "iv"],
"filter_type": ["top_k", "threshold"],
"take_high": [True, True],
"threshold": [10, 0.001]
},
"select_col_indexes": -1
}
hetero_feature_selection_0 = HeteroFeatureSelection(**param)
param = {
"k": 3,
"max_iter": 10
}
hetero_kmeans_0 = HeteroKmeans(name='hetero_kmeans_0', **param)
evaluation_0 = Evaluation(name='evaluation_0', eval_type='clustering')
# add components to pipeline, in order of task execution
pipeline.add_component(reader_0)
pipeline.add_component(data_transform_0, data=Data(data=reader_0.output.data))
# set data input sources of intersection components
pipeline.add_component(intersection_0, data=Data(data=data_transform_0.output.data))
# set train & validate data of hetero_lr_0 component
pipeline.add_component(hetero_feature_binning_0, data=Data(data=intersection_0.output.data))
pipeline.add_component(hetero_feature_selection_0, data=Data(data=intersection_0.output.data),
model=Model(isometric_model=hetero_feature_binning_0.output.model))
pipeline.add_component(hetero_kmeans_0, data=Data(train_data=hetero_feature_selection_0.output.data))
print(f"data: {hetero_kmeans_0.output.data.data[0]}")
pipeline.add_component(evaluation_0, data=Data(data=hetero_kmeans_0.output.data.data[0]))
# compile pipeline once finished adding modules, this step will form conf and dsl files for running job
pipeline.compile()
# fit model
pipeline.fit()
# query component summary
print(pipeline.get_component("hetero_kmeans_0").get_summary())
if __name__ == "__main__":
parser = argparse.ArgumentParser("PIPELINE DEMO")
parser.add_argument("-config", type=str,
help="config file")
args = parser.parse_args()
if args.config is not None:
main(args.config)
else:
main()
init.py
pipeline-kmeans-validate.py
import argparse
from pipeline.backend.pipeline import PipeLine
from pipeline.component import DataTransform
from pipeline.component import HeteroKmeans
from pipeline.component import Intersection
from pipeline.component import Evaluation
from pipeline.component import Reader
from pipeline.interface import Data
from pipeline.interface import Model
from pipeline.utils.tools import load_job_config
def main(config="../../config.yaml", namespace=""):
# obtain config
if isinstance(config, str):
config = load_job_config(config)
parties = config.parties
guest = parties.guest[0]
host = parties.host[0]
arbiter = parties.arbiter[0]
guest_train_data = {"name": "breast_hetero_guest", "namespace": f"experiment{namespace}"}
host_train_data = {"name": "breast_hetero_host", "namespace": f"experiment{namespace}"}
guest_eval_data = {"name": "breast_hetero_guest", "namespace": f"experiment{namespace}"}
host_eval_data = {"name": "breast_hetero_host", "namespace": f"experiment{namespace}"}
# initialize pipeline
pipeline = PipeLine()
# set job initiator
pipeline.set_initiator(role='guest', party_id=guest)
# set participants information
pipeline.set_roles(guest=guest, host=host, arbiter=arbiter)
# define Reader components to read in data
reader_0 = Reader(name="reader_0")
# configure Reader for guest
reader_0.get_party_instance(role='guest', party_id=guest).component_param(table=guest_train_data)
# configure Reader for host
reader_0.get_party_instance(role='host', party_id=host).component_param(table=host_train_data)
reader_1 = Reader(name="reader_1")
reader_1.get_party_instance(role='guest', party_id=guest).component_param(table=guest_eval_data)
reader_1.get_party_instance(role='host', party_id=host).component_param(table=host_eval_data)
# define DataTransform components
data_transform_0 = DataTransform(name="data_transform_0") # start component numbering at 0
data_transform_1 = DataTransform(name="data_transform_1")
# get DataTransform party instance of guest
data_transform_0_guest_party_instance = data_transform_0.get_party_instance(role='guest', party_id=guest)
# configure DataTransform for guest
data_transform_0_guest_party_instance.component_param(with_label=True, output_format="dense")
# get and configure DataTransform party instance of host
data_transform_0.get_party_instance(role='host', party_id=host).component_param(with_label=False)
# define Intersection components
intersection_0 = Intersection(name="intersection_0")
intersection_1 = Intersection(name="intersection_1")
param = {
"k": 3,
"max_iter": 10
}
hetero_kmeans_0 = HeteroKmeans(name='hetero_kmeans_0', **param)
hetero_kmeans_1 = HeteroKmeans(name='hetero_kmeans_1')
evaluation_0 = Evaluation(name='evaluation_0', eval_type='clustering')
evaluation_1 = Evaluation(name='evaluation_1', eval_type='clustering')
# add components to pipeline, in order of task execution
pipeline.add_component(reader_0)
pipeline.add_component(reader_1)
pipeline.add_component(data_transform_0, data=Data(data=reader_0.output.data))
pipeline.add_component(
data_transform_1, data=Data(
data=reader_1.output.data), model=Model(
data_transform_0.output.model))
# set data input sources of intersection components
pipeline.add_component(intersection_0, data=Data(data=data_transform_0.output.data))
pipeline.add_component(intersection_1, data=Data(data=data_transform_1.output.data))
# set train & validate data of hetero_lr_0 component
pipeline.add_component(hetero_kmeans_0, data=Data(train_data=intersection_0.output.data))
pipeline.add_component(hetero_kmeans_1, data=Data(train_data=intersection_1.output.data))
# print(f"data: {hetero_kmeans_0.output.data.data[0]}")
pipeline.add_component(evaluation_0, data=Data(data=hetero_kmeans_0.output.data.data[0]))
pipeline.add_component(evaluation_1, data=Data(data=hetero_kmeans_1.output.data.data[0]))
# compile pipeline once finished adding modules, this step will form conf and dsl files for running job
pipeline.compile()
# fit model
pipeline.fit()
# query component summary
print(pipeline.get_component("hetero_kmeans_0").get_summary())
if __name__ == "__main__":
parser = argparse.ArgumentParser("PIPELINE DEMO")
parser.add_argument("-config", type=str,
help="config file")
args = parser.parse_args()
if args.config is not None:
main(args.config)
else:
main()
## Hetero Kmeans Configuration Usage Guide.
#### Example Tasks
This section introduces the dsl and conf for different types of tasks.
1. Train Task:
dsl: test_hetero_kmeans_train_dsl.json
runtime_config : test_hetero_kmeans_train_conf.json
2. Validate Task (with early-stopping parameters specified):
dsl: test_hetero_kmeans_validate_dsl.json
runtime_config : test_hetero_kmeans_validate_conf.json
3. Multi-host Train Task:
dsl: test_hetero_kmeans_multi_host_dsl.json
conf: test_hetero_kmeans_multi_host_conf.json
4. With Feature-engineering Task:
dsl: test_hetero_kmeans_with_feature_engineering_dsl.json
conf: test_hetero_kmeans_with_feature_engineering_conf.json
Users can use following commands to run a task.
bash flow job submit -c ${runtime_config} -d ${dsl}
hetero_kmeans_testsuite.json
{
"data": [
{
"file": "examples/data/breast_hetero_guest.csv",
"head": 1,
"partition": 16,
"table_name": "breast_hetero_guest",
"namespace": "experiment",
"role": "guest_0"
},
{
"file": "examples/data/breast_hetero_host.csv",
"head": 1,
"partition": 16,
"table_name": "breast_hetero_host",
"namespace": "experiment",
"role": "host_0"
},
{
"file": "examples/data/breast_hetero_host.csv",
"head": 1,
"partition": 16,
"table_name": "breast_hetero_host",
"namespace": "experiment",
"role": "host_1"
},
{
"file": "examples/data/vehicle_scale_hetero_guest.csv",
"head": 1,
"partition": 16,
"table_name": "vehicle_scale_hetero_guest",
"namespace": "experiment",
"role": "guest_0"
},
{
"file": "examples/data/vehicle_scale_hetero_host.csv",
"head": 1,
"partition": 16,
"table_name": "vehicle_scale_hetero_host",
"namespace": "experiment",
"role": "host_0"
}
],
"tasks": {
"kmeans": {
"conf": "test_hetero_kmeans_conf.json",
"dsl": "test_hetero_kmeans_dsl.json"
},
"kmeans_validate": {
"conf": "test_hetero_kmeans_validate_conf.json",
"dsl": "test_hetero_kmeans_validate_dsl.json"
},
"kmeans_feature_engineering": {
"conf": "test_hetero_kmeans_with_feature_engineering_conf.json",
"dsl": "test_hetero_kmeans_with_feature_engineering_dsl.json"
},
"kmeans_multi_host": {
"conf": "test_hetero_kmeans_multi_host_conf.json",
"dsl": "test_hetero_kmeans_multi_host_dsl.json"
}
}
}
test_hetero_kmeans_validate_conf.json
{
"dsl_version": 2,
"initiator": {
"role": "guest",
"party_id": 9999
},
"role": {
"arbiter": [
9999
],
"host": [
9998
],
"guest": [
9999
]
},
"component_parameters": {
"common": {
"hetero_kmeans_0": {
"k": 3,
"max_iter": 10
},
"evaluation_0": {
"eval_type": "clustering"
},
"evaluation_1": {
"eval_type": "clustering"
}
},
"role": {
"host": {
"0": {
"data_transform_0": {
"with_label": false
},
"reader_1": {
"table": {
"name": "breast_hetero_host",
"namespace": "experiment"
}
},
"reader_0": {
"table": {
"name": "breast_hetero_host",
"namespace": "experiment"
}
}
}
},
"guest": {
"0": {
"data_transform_0": {
"with_label": true,
"output_format": "dense"
},
"reader_1": {
"table": {
"name": "breast_hetero_guest",
"namespace": "experiment"
}
},
"reader_0": {
"table": {
"name": "breast_hetero_guest",
"namespace": "experiment"
}
}
}
}
}
}
}
test_hetero_kmeans_dsl.json
{
"components": {
"reader_0": {
"module": "Reader",
"output": {
"data": [
"data"
]
}
},
"data_transform_0": {
"module": "DataTransform",
"input": {
"data": {
"data": [
"reader_0.data"
]
}
},
"output": {
"data": [
"data"
],
"model": [
"model"
]
}
},
"intersection_0": {
"module": "Intersection",
"input": {
"data": {
"data": [
"data_transform_0.data"
]
}
},
"output": {
"data": [
"data"
]
}
},
"hetero_kmeans_0": {
"module": "HeteroKmeans",
"input": {
"data": {
"train_data": [
"intersection_0.data"
]
}
},
"output": {
"data": [
"data_0",
"data_1"
],
"model": [
"model"
]
}
},
"evaluation_0": {
"module": "Evaluation",
"input": {
"data": {
"data": [
"hetero_kmeans_0.data_0"
]
}
},
"output": {
"data": [
"data"
]
}
}
}
}
test_hetero_kmeans_conf.json
{
"dsl_version": 2,
"initiator": {
"role": "guest",
"party_id": 9999
},
"role": {
"arbiter": [
9999
],
"host": [
9998
],
"guest": [
9999
]
},
"component_parameters": {
"common": {
"hetero_kmeans_0": {
"k": 3,
"max_iter": 10
},
"evaluation_0": {
"eval_type": "clustering"
}
},
"role": {
"host": {
"0": {
"data_transform_0": {
"with_label": false
},
"reader_0": {
"table": {
"name": "breast_hetero_host",
"namespace": "experiment"
}
}
}
},
"guest": {
"0": {
"data_transform_0": {
"with_label": true,
"output_format": "dense"
},
"reader_0": {
"table": {
"name": "breast_hetero_guest",
"namespace": "experiment"
}
}
}
}
}
}
}
test_hetero_kmeans_validate_dsl.json
{
"components": {
"reader_0": {
"module": "Reader",
"output": {
"data": [
"data"
]
}
},
"reader_1": {
"module": "Reader",
"output": {
"data": [
"data"
]
}
},
"data_transform_0": {
"module": "DataTransform",
"input": {
"data": {
"data": [
"reader_0.data"
]
}
},
"output": {
"data": [
"data"
],
"model": [
"model"
]
}
},
"data_transform_1": {
"module": "DataTransform",
"input": {
"data": {
"data": [
"reader_1.data"
]
},
"model": [
"data_transform_0.model"
]
},
"output": {
"data": [
"data"
],
"model": [
"model"
]
}
},
"intersection_0": {
"module": "Intersection",
"input": {
"data": {
"data": [
"data_transform_0.data"
]
}
},
"output": {
"data": [
"data"
]
}
},
"intersection_1": {
"module": "Intersection",
"input": {
"data": {
"data": [
"data_transform_1.data"
]
}
},
"output": {
"data": [
"data"
]
}
},
"hetero_kmeans_0": {
"module": "HeteroKmeans",
"input": {
"data": {
"train_data": [
"intersection_0.data"
]
}
},
"output": {
"data": [
"data_0",
"data_1"
],
"model": [
"model"
]
}
},
"hetero_kmeans_1": {
"module": "HeteroKmeans",
"input": {
"data": {
"train_data": [
"intersection_1.data"
]
}
},
"output": {
"data": [
"data_0",
"data_1"
],
"model": [
"model"
]
}
},
"evaluation_0": {
"module": "Evaluation",
"input": {
"data": {
"data": [
"hetero_kmeans_0.data_0"
]
}
},
"output": {
"data": [
"data"
]
}
},
"evaluation_1": {
"module": "Evaluation",
"input": {
"data": {
"data": [
"hetero_kmeans_1.data_0"
]
}
},
"output": {
"data": [
"data"
]
}
}
}
}
test_hetero_kmeans_with_feature_engineering_dsl.json
{
"components": {
"reader_0": {
"module": "Reader",
"output": {
"data": [
"data"
]
}
},
"data_transform_0": {
"module": "DataTransform",
"input": {
"data": {
"data": [
"reader_0.data"
]
}
},
"output": {
"data": [
"data"
],
"model": [
"model"
]
}
},
"intersection_0": {
"module": "Intersection",
"input": {
"data": {
"data": [
"data_transform_0.data"
]
}
},
"output": {
"data": [
"data"
]
}
},
"hetero_feature_binning_0": {
"module": "HeteroFeatureBinning",
"input": {
"data": {
"data": [
"intersection_0.data"
]
}
},
"output": {
"data": [
"data"
],
"model": [
"model"
]
}
},
"hetero_feature_selection_0": {
"module": "HeteroFeatureSelection",
"input": {
"data": {
"data": [
"intersection_0.data"
]
},
"isometric_model": [
"hetero_feature_binning_0.model"
]
},
"output": {
"data": [
"data"
],
"model": [
"model"
]
}
},
"hetero_kmeans_0": {
"module": "HeteroKmeans",
"input": {
"data": {
"train_data": [
"hetero_feature_selection_0.data"
]
}
},
"output": {
"data": [
"data_0",
"data_1"
],
"model": [
"model"
]
}
},
"evaluation_0": {
"module": "Evaluation",
"input": {
"data": {
"data": [
"hetero_kmeans_0.data_0"
]
}
},
"output": {
"data": [
"data"
]
}
}
}
}
test_hetero_kmeans_multi_host_conf.json
{
"dsl_version": 2,
"initiator": {
"role": "guest",
"party_id": 9999
},
"role": {
"arbiter": [
9999
],
"host": [
9998,
10000
],
"guest": [
9999
]
},
"component_parameters": {
"common": {
"hetero_kmeans_0": {
"k": 3,
"max_iter": 10
},
"evaluation_0": {
"eval_type": "clustering"
}
},
"role": {
"host": {
"1": {
"reader_0": {
"table": {
"name": "breast_hetero_host",
"namespace": "experiment"
}
},
"reader_1": {
"table": {
"name": "breast_hetero_host",
"namespace": "experiment"
}
},
"data_transform_0": {
"with_label": false
}
},
"0": {
"reader_0": {
"table": {
"name": "breast_hetero_host",
"namespace": "experiment"
}
},
"reader_1": {
"table": {
"name": "breast_hetero_host",
"namespace": "experiment"
}
},
"data_transform_0": {
"with_label": false
}
}
},
"guest": {
"0": {
"reader_0": {
"table": {
"name": "breast_hetero_guest",
"namespace": "experiment"
}
},
"reader_1": {
"table": {
"name": "breast_hetero_guest",
"namespace": "experiment"
}
},
"data_transform_0": {
"with_label": true,
"output_format": "dense"
}
}
}
}
}
}
test_hetero_kmeans_multi_host_dsl.json
{
"components": {
"reader_0": {
"module": "Reader",
"output": {
"data": [
"data"
]
}
},
"data_transform_0": {
"module": "DataTransform",
"input": {
"data": {
"data": [
"reader_0.data"
]
}
},
"output": {
"data": [
"data"
],
"model": [
"model"
]
}
},
"intersection_0": {
"module": "Intersection",
"input": {
"data": {
"data": [
"data_transform_0.data"
]
}
},
"output": {
"data": [
"data"
]
}
},
"hetero_kmeans_0": {
"module": "HeteroKmeans",
"input": {
"data": {
"train_data": [
"intersection_0.data"
]
}
},
"output": {
"data": [
"data_0",
"data_1"
],
"model": [
"model"
]
}
},
"evaluation_0": {
"module": "Evaluation",
"input": {
"data": {
"data": [
"hetero_kmeans_0.data_0"
]
}
},
"output": {
"data": [
"data"
]
}
}
}
}
test_hetero_kmeans_with_feature_engineering_conf.json
{
"dsl_version": 2,
"initiator": {
"role": "guest",
"party_id": 9999
},
"role": {
"arbiter": [
9999
],
"host": [
9998
],
"guest": [
9999
]
},
"component_parameters": {
"common": {
"hetero_feature_binning_0": {
"method": "optimal",
"bin_indexes": -1,
"optimal_binning_param": {
"metric_method": "iv"
}
},
"hetero_feature_selection_0": {
"select_col_indexes": -1,
"filter_methods": [
"manually",
"iv_filter"
],
"manually_param": {
"filter_out_indexes": [
1
]
},
"iv_param": {
"metrics": [
"iv",
"iv"
],
"filter_type": [
"top_k",
"threshold"
],
"take_high": [
true,
true
],
"threshold": [
10,
0.001
]
}
},
"hetero_kmeans_0": {
"k": 3,
"max_iter": 10
},
"evaluation_0": {
"eval_type": "clustering"
}
},
"role": {
"host": {
"0": {
"data_transform_0": {
"with_label": false
},
"reader_0": {
"table": {
"name": "breast_hetero_host",
"namespace": "experiment"
}
}
}
},
"guest": {
"0": {
"data_transform_0": {
"with_label": true,
"output_format": "dense"
},
"reader_0": {
"table": {
"name": "breast_hetero_guest",
"namespace": "experiment"
}
}
}
}
}
}
}